SNP Testing Services Overview
Single nucleotide polymorphism (SNP) testing is used for annual monitoring of the genetic background of rats and mice, detection of gene mutations or genetic contaminations and batch testing of rat and mouse mutants.
Why GemPharmatech
One-stop solution: We provide a one-stop solution for SNP testing from primer design to result reporting.
Rich experience: Since establishing the high-throughput testing platform, we have cumulatively tested more than 100,000 data points.
Stable genetic testing panel: We have successfully developed SNP panels applicable to genetic testing and speed congenics for 13 inbred mouse strains.
SNP Testing Services
SNP testing refers to the conversion, transversion, insertion, and deletion of a single base arising from the mutation of a single nucleotide with a mutation frequency greater than 1% in a population. As a third-generation genetic molecular marker, the SNP labeling technique possesses numerous advantages in genetic quality monitoring as compared with other genetic markers, including wide distribution, good genetic stability, impressive representativeness, and easier implementation of high-throughput and automatic monitoring. As a result, it has been widely applied in agriculture, pharmaceutical, life science, and other research fields.
We can perform SNP testing using KASP (Kompetitive Allele Specific PCR) for the following purposes:
Annual monitoring of genetic background of rats and mice: Annual genetic testing for laboratory animals is an essential approach to evaluate and ensure the quality of laboratory animals, especially for which regular genetic background monitoring is necessary.
Testing for genetic mutation and contamination in rats and mice: This test should be performed when a new phenotype is identified or when a mismatched rat/mouse is suspected during a study.
Batch testing of rat and mouse mutants: SNP testing can be used for precise genotyping of InDels (insertions and deletions).
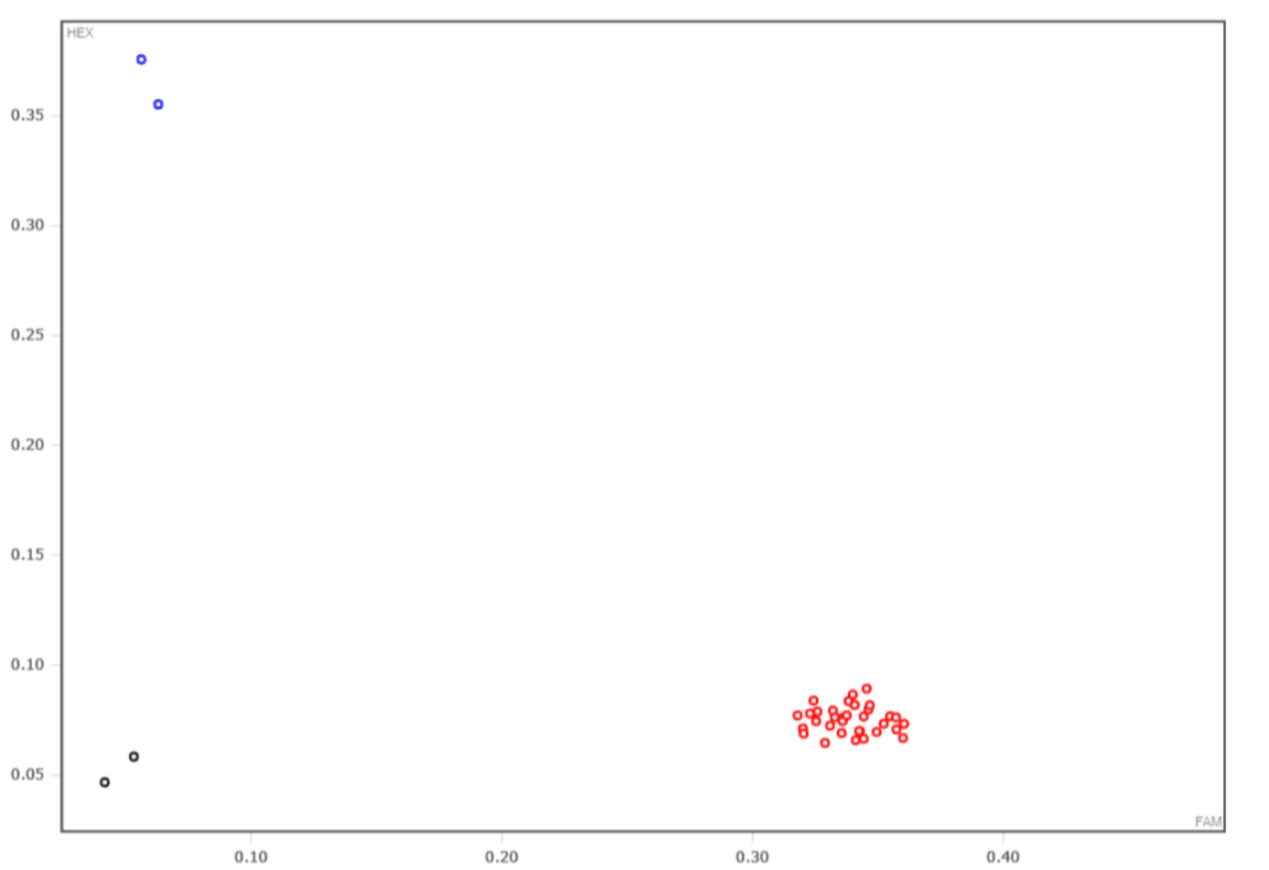
X: HEX Channel Y: FAM Channel
Red dots:FAM/FAM, Blue dots: HEX/HEX, Black dots: blank control.
High-throughput KASP based on available SNP loci information is utilized for precise allele genotyping of specified SNPs and InDels in diverse genome samples (including complicated genomic DNA samples), employing fluorescence detection with high sensitivity. In terms of technological principle, it is identical to Taqman assay (the gold standard) and produces consistent accuracy. Unlike traditional Taqman assays, however, amplification of the general fluorescence primer is available for assay at all loci based on the unique ARM PCR approach, rather than requiring a probe or primer generated for each target. We have optimized PCR methods to address the needs of high-throughput reactions at various loci. While the accuracy of KASP is comparable to the gold standard Taqman test, it is more economical and has better loci adaptability.

